
SCIENTIFIC
The Science Behind The 4Kscore® Test |
The 4Kscore® Test is a follow-up blood test after an abnormal PSA (Prostate Specific Antigen) or digital rectal exam (DRE) to provide the probability of finding aggressive prostate cancer if a biopsy were to be performed. The 4Kscore® Test combines measurements of four prostate-specific biomarkers along with patient clinical information in an algorithm to accurately predict a man’s pobability for aggressive prostate cancer. Understanding the probability of finding aggressive prostate cancer gives a patient and a physician more information to decide if a prostate biopsy is warranted.
The 4Kscore® Biomarkers
The 4Kscore® Test measures four prostate-specific kallikreins: Total PSA, Free PSA, Intact PSA, and human kallikrein 2 (hk2) in addition to incorporating clinical results. Total PSA and Free PSA are important markers for prostate cancer but have low specificity for risk of aggressive prostate cancer. Intact PSA and human kallikrein 2 (hk2) are present in much lower concentrations than total PSA and Free PSA, and changes in these biomarkers are associated with aggressive types of prostate cancer.
A prospective U.S.–based validation study of The 4Kscore® Test in 1012 men examined the accuracy of the test to predict the probability of high-grade cancer on biopsy and the contributions of the different test components. The Area Under the Curve (AUC) for The 4Kscore® Test was 0.82 compared to 0.74 for a PSA–based probablity calculator. Intact PSA and human kallikrein 2 (hk2) contributed 0.07 to the AUC of The 4Kscore Test, which was a significant contribution. The 4Kscore® Test predicted probability of high-grade cancer with high calibration to the true probability observed in the study.
The 4Kscore® Score Test Is FDA Approved, and Its Accuracy Has Been Demonstrated in Two Multi-Center Prospective Validation Studies in the United States
Parekh DJ et al. Eur Urol. 2015 Sep;68(3):464-70
- 1012 men prospectively enrolled in 26 academic and community practices
- Subjects had scheduled prostate biopsy as recommended by a urologist and blood drawn prior to biopsy
- Minimum of 10 core TRUS-guided biopsies performed on all patients
- AUC = 0.82
- 36% of biopsies could be avoided or delayed
A Multi-Institutional Prospective Trial Confirms Noninvasive Blood Test Maintains Predictive Value in African American Men
Study Results Show The 4Kscore® Test Performance at 5.0 Cut Point in the African American Population Identifies Men with High-Grade Prostate Cancer with 99% Sensitivity and 95% Negative Predictive Value (The 4Kscore® Test PI)
Punnen S et al. J Urol. 2018 Jun;199(6):1459-1463:
- 8 Veterans Affairs Hospitals
- 56% of the 366 men were African American
- Similar 4Kscore® performance to first validation study
- No significant difference in 4Kscore® accuracy between African American and non-African American men
- Sensitivity of 97% and Negative Predictive Value (NPV) of 95% for the detection of aggressive prostate cancer
- 22% of study subjects could avoid or delay biopsies
Significant Clinical Research Findings Supporting 4Kscore®
Distant Prostate Cancer Metastases After 20 Years Stratified by 4Kscore®
Improving the Specificity of Screening for Lethal Prostate Cancer Using Prostate-specific Antigen and a Panel of Kallikrein Markers: A Nested Case–Control Study
Stattin et al. Eur Urol. 2015;68(2):207-213:
- Followed 12,542 men enrolled in Västerbotten Intervention Project from 1986-2009
- Enrollees received very limited PSA testing for next 20 years
- After 20 years, the development of distant prostate cancer metastases was determined from records in the Swedish Cancer Registry
- A single blood sample obtained at enrollment was tested for PSA
- Men ages 50 to 60 at enrollment were further tested with The 4Kscore® Test if the PSA was elevated
In 50- and 60-year-old men with an elevated PSA, a low-risk 4Kscore® was shown to predict low probability for metastasis.
Prostate Cancer Mortality After 20 Years Predicted by The 4Kscore® Test
“Twenty-year Risk of Prostate Cancer Death by Midlife Prostate-specific Antigen and a Panel of Four Kallikrein Markers in a Large Population-based Cohort of Healthy Men”
Sjoberg et al. Eur Urol. 2018;73(6):941-948:
- Followed 11,506 healthy men ages 45-73 enrolled in Malmö Diet and Cancer Study from 1991-1996
- Very limited PSA testing for subsequent 20 years
- A blood sample was cryopreserved and the men were followed through December 2014 when the blood samples were tested
- After 20 years, prostate cancer mortality was determined from records in the Swedish Cancer Registry
- A single blood sample obtained at enrollment was tested for PSA
- 317 deaths from prostate cancer were found in the cohort
- Men with elevated PSA were further tested with 4Kscore®
The 4Kscore® Test can stratify men and predict distant prostate cancer mortality.
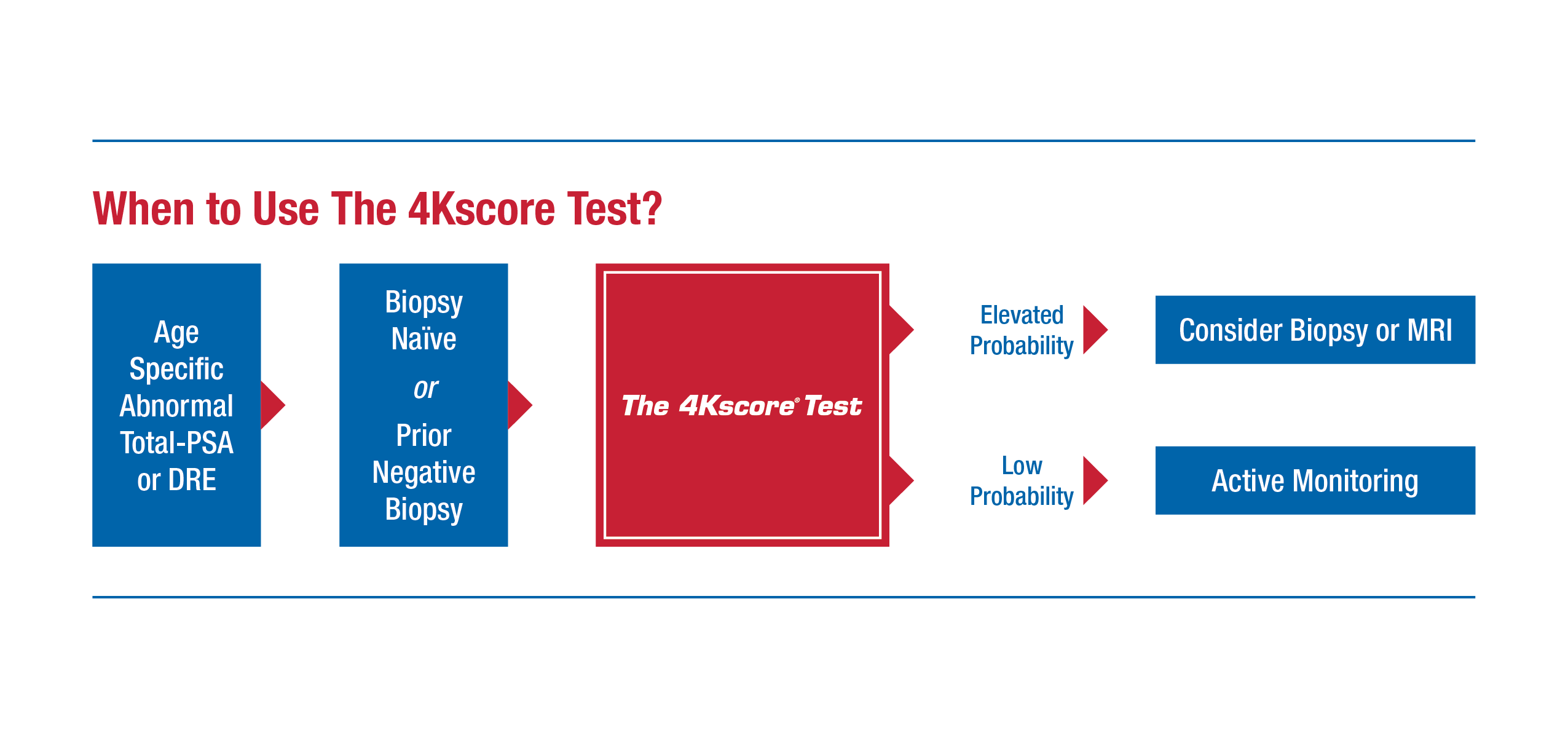
The 4Kscore® Test: A Precision Approach to Prostate Cancer Evaluation
- PSA is an effective screening test for prostate cancer, but it lacks specificity for the aggressive form of prostate cancer
- The 4Kscore® Test offers the sensitivity and specificity needed to detect aggressive prostate cancer probability
- The 4Kscore® Test is clinically validated in two U.S.–based, prospective studies, including a 2017 study where the majority of subjects were African American
- When probability score of 7.5% is used for assessment, no Gleason ≥8 was missed
- Significant number of study subjects could have avoided biopsies
- Research has demonstrated that The 4Kscore® Test can probability-stratify men for prostate cancer metastasis and mortality for up to 20 years
- Multiple peer-reviewed publications, with studies involving more than 20,000 men, demonstrate the clinical effectiveness of The 4Kscore® Test
- FDA Approved
- Included in current U.S., Canadian, and European prostate cancer early-detection guidelines
- CPT Category I code (81539)
- The CPT code provided is based on guidance from the American Medical Association (AMA) and is for informational purposes only
- CPT coding is the sole responsibility of the billing party
References
Parekh et al. A multi-institutional prospective trial in the USA confirms that the 4Kscore® accurately identifies men with high-grade prostate cancer. Eur Urol. 2015;68(3):464-470.
Punnen et al. A multi-institutional prospective trial confirms noninvasive blood test maintains predictive value in African American men. J Urol. 2018;199(6);1459-1463.
Stattin et al. Improving the Specificity of Screening for Lethal Prostate cancer using prostate-specific antigen and a panel of kallikrein markers: a nested case-control study. Eur Urol. 2015;68(2):207-213.
Sjoberg et al. Twenty-year risk of prostate cancer death by midlife prostate-specific antigen and a panel of four kallikrein markers in a large population-based cohort of healthy men. Eur Urol. 2018;73(6):941-948.
Peltola MT, Niemelä P, Väisänen V, et al. Intact and internally cleaved free prostate‐specific antigen in patients with prostate cancer with different pathologic stages and grades. Urology 2011 Apr;77(4):1009.e1.
Steuber T, Vickers AJ, Serio AM, et al. Comparison of free and total forms of serum human kallikrein 2 and prostate‐specific antigen for prediction of locally advanced and recurrent prostate cancer. Clin Chem. 2007 Feb;53(2):233‐40.
Wenske S, Korets R, Cronin AM, et al. Evaluation of molecular forms of prostate-specific antigen and human kallikrein 2 in predicting biochemical failure after radical prostatectomy. Int J Cancer. 2009 Feb 1;124(3):659‐653.
Find The 4Kscore® Test Near You
You can find a Patient Service Center by using the link below.
Este sitio web (y las páginas contenidas) estan desactivado. Si se necesita asistencia en español u otro idioma, por favor llame sin cargas a 800 229 5227.
close
